import sympy as sp
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import Math, display
from symbolic_fem_workbench import (
build_poisson_triangle_p1_local_problem,
assemble_dense_matrix, assemble_dense_vector,
apply_dirichlet_by_reduction, expand_reduced_solution,
)8 Manual 2D Assembly: From Local Tensors to Global Solution
This notebook demonstrates the assembly process: combining element matrices and vectors into a global system, applying boundary conditions, and solving for the finite element solution. Assembly is fundamentally a topological operation—it depends only on node connectivity, not on the analytical properties of elements.
8.1 Step 1: Get symbolic local tensors and convert to numerical
We retrieve the element stiffness matrix and load vector from the previous notebook and convert them to numerical functions via lambdify.
data = build_poisson_triangle_p1_local_problem()
geom = data["geometry"]
ke_fn = sp.lambdify(
(geom.x1, geom.y1, geom.x2, geom.y2, geom.x3, geom.y3),
data["Ke"], "numpy"
)
fe_fn = sp.lambdify(
(geom.x1, geom.y1, geom.x2, geom.y2, geom.x3, geom.y3, data["f"]),
data["fe"], "numpy"
)
print("Numerical kernels ready.")Numerical kernels ready.8.2 Step 2: Define a tiny mesh
We discretize the unit square using 4 triangles arranged around a central point. This simple mesh illustrates the assembly process without excessive complexity.
nodes = np.array([
[0.0, 0.0], # 0: bottom-left
[1.0, 0.0], # 1: bottom-right
[1.0, 1.0], # 2: top-right
[0.0, 1.0], # 3: top-left
[0.5, 0.5], # 4: center
])
elements = [(0,1,4), (1,2,4), (2,3,4), (3,0,4)]
# Visualize the mesh
fig, ax = plt.subplots(1, 1, figsize=(5, 5))
for tri in elements:
pts = nodes[list(tri) + [tri[0]]]
ax.plot(pts[:, 0], pts[:, 1], "b-")
for i, (x, y) in enumerate(nodes):
ax.plot(x, y, "ko", markersize=8)
ax.annotate(f" {i}", (x, y), fontsize=12)
ax.set_title("Mesh: 4 triangles, 5 nodes")
ax.set_aspect("equal")
plt.show()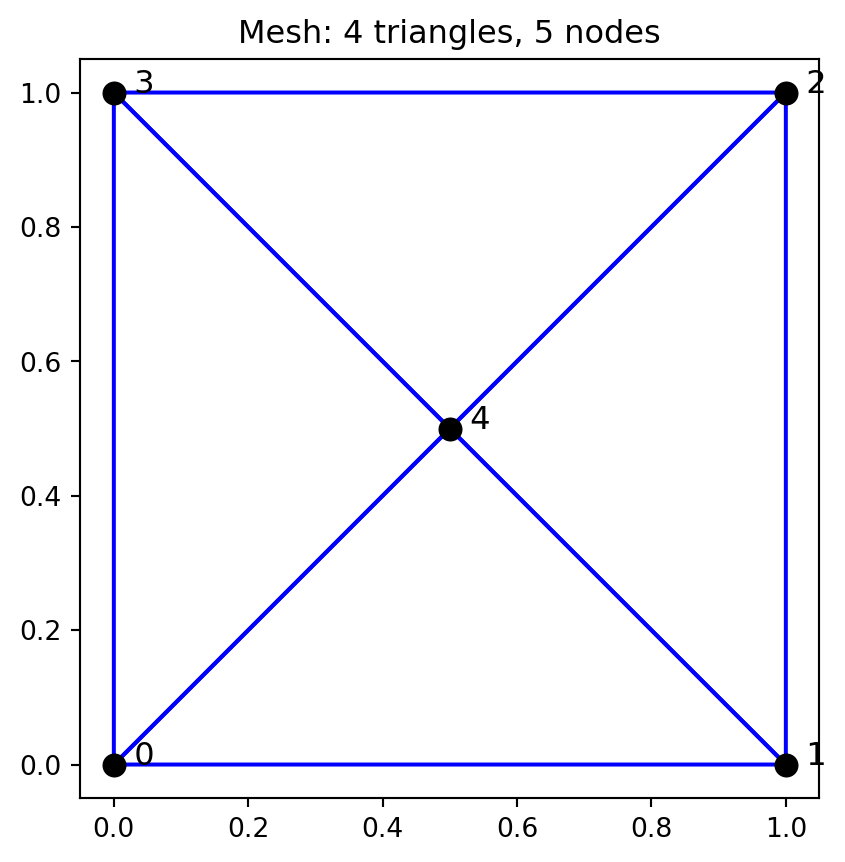
8.3 Step 3: Element-by-element assembly
The assembly loop scans through all elements, retrieves element matrices/vectors, and accumulates them into global arrays using the connectivity information.
ndofs = len(nodes)
K_global = np.zeros((ndofs, ndofs))
F_global = np.zeros(ndofs)
source_value = 1.0
for eid, conn in enumerate(elements):
coords = nodes[list(conn)]
x1v, y1v = coords[0]
x2v, y2v = coords[1]
x3v, y3v = coords[2]
K_local = np.asarray(ke_fn(x1v, y1v, x2v, y2v, x3v, y3v), dtype=float)
F_local = np.asarray(fe_fn(x1v, y1v, x2v, y2v, x3v, y3v, source_value), dtype=float).reshape(-1)
print(f"Element {eid}, connectivity {conn}")
print(f" K_local diagonal: {np.diag(K_local)}")
print(f" F_local: {F_local}")
assemble_dense_matrix(K_global, K_local, conn)
assemble_dense_vector(F_global, F_local, conn)
print("\nAssembled K_global:")
print(np.round(K_global, 4))
print("\nAssembled F_global:")
print(np.round(F_global, 4))Element 0, connectivity (0, 1, 4)
K_local diagonal: [0.5 0.5 1. ]
F_local: [0.08333333 0.08333333 0.08333333]
Element 1, connectivity (1, 2, 4)
K_local diagonal: [0.5 0.5 1. ]
F_local: [0.08333333 0.08333333 0.08333333]
Element 2, connectivity (2, 3, 4)
K_local diagonal: [0.5 0.5 1. ]
F_local: [0.08333333 0.08333333 0.08333333]
Element 3, connectivity (3, 0, 4)
K_local diagonal: [0.5 0.5 1. ]
F_local: [0.08333333 0.08333333 0.08333333]
Assembled K_global:
[[ 1. 0. 0. 0. -1.]
[ 0. 1. 0. 0. -1.]
[ 0. 0. 1. 0. -1.]
[ 0. 0. 0. 1. -1.]
[-1. -1. -1. -1. 4.]]
Assembled F_global:
[0.1667 0.1667 0.1667 0.1667 0.3333]8.4 Step 4: Apply Dirichlet BCs and solve
Dirichlet boundary conditions are applied by reducing the system: we eliminate rows and columns corresponding to constrained DOFs and solve for the free DOFs, then expand the solution.
constrained = [0, 1, 2, 3] # All boundary nodes
K_ff, F_f, free_dofs = apply_dirichlet_by_reduction(K_global, F_global, constrained)
print(f"Free DOFs: {free_dofs}")
print(f"K_ff = {K_ff}")
print(f"F_f = {F_f}")
u_free = np.linalg.solve(K_ff, F_f)
u = expand_reduced_solution(u_free, ndofs, free_dofs, constrained)
print(f"\nSolution vector u = {u}")
print(f"Center node value u[4] = {u[4]:.6f}")
print(f"Expected (analytical): 1/12 = {1/12:.6f}")
print(f"Match: {np.isclose(u[4], 1/12)}")Free DOFs: [4]
K_ff = [[4.]]
F_f = [0.33333333]
Solution vector u = [0. 0. 0. 0. 0.08333333]
Center node value u[4] = 0.083333
Expected (analytical): 1/12 = 0.083333
Match: True8.5 Step 5: Visualize the solution
We plot the FEM solution as a colored patch over the mesh, with annotations showing nodal values.
from matplotlib.tri import Triangulation
tri_conn = np.array(elements)
triang = Triangulation(nodes[:, 0], nodes[:, 1], tri_conn)
fig, ax = plt.subplots(figsize=(6, 5))
tpc = ax.tripcolor(triang, u, shading="flat", cmap="viridis")
ax.triplot(triang, "k-", linewidth=0.5)
for i, (x, y) in enumerate(nodes):
ax.annotate(f"{i}: u={u[i]:.4f}", (x, y), fontsize=9,
xytext=(5, 5), textcoords="offset points")
plt.colorbar(tpc, label="u")
ax.set_title("FEM Solution: Poisson on 4-Triangle Mesh")
ax.set_aspect("equal")
plt.show()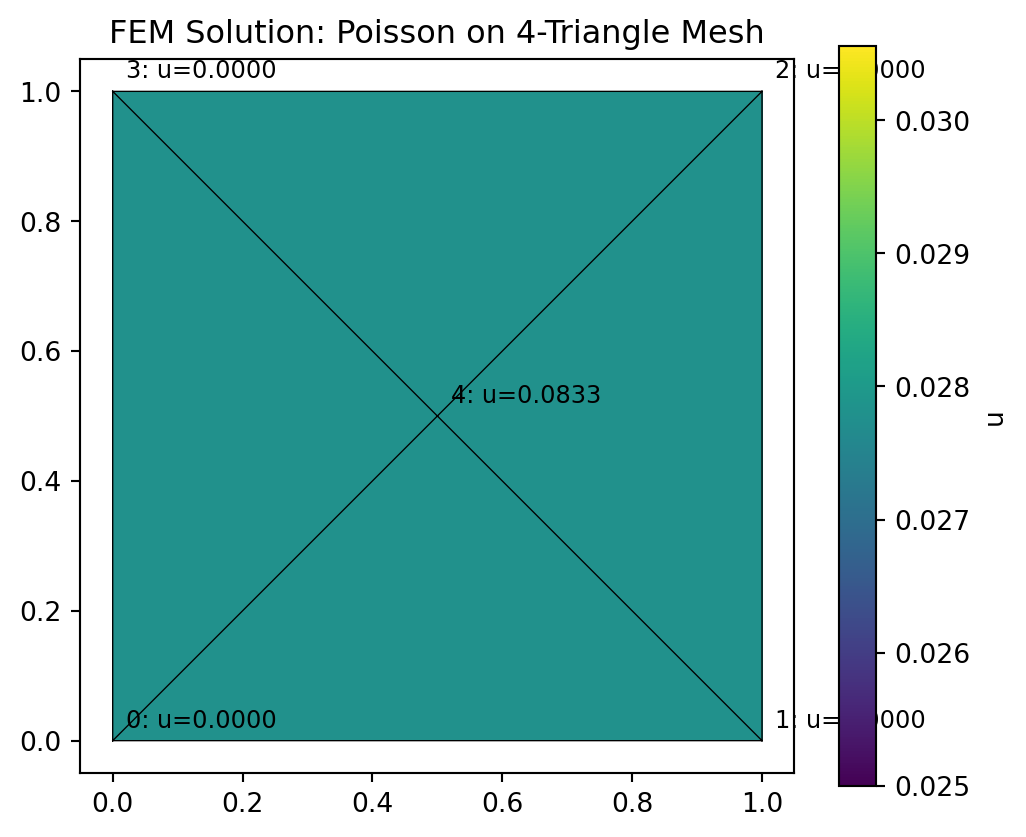
8.6 Exercise
Question: Move the center node from \((0.5, 0.5)\) to \((0.3, 0.7)\). How does the solution change? Is the center node value still \(1/12\)? Why or why not?
Hint: The center node is no longer at a symmetric position, so the Jacobians of the four elements are all different. This affects their stiffness matrices.
# Exercise: Move the center node from (0.5, 0.5) to (0.3, 0.7).
# How does the solution change? Is the center node value still 1/12?
# Why or why not?